|
Regulatory proteins (RPs) are involved in the control of diverse cellular systems, allowing adaptive responses to changes in environmental conditions. They are found in prokaryotes where they control diverse aspects of prokaryotic metabolism, such as cell differentiation, morphogenesis, central metabolism, motility, biofilm formation and virulence. To understand RP functions, it is critical to characterise and analyse their modular complexity. P2RP, a freely accessible web server, has been developed for computational analysis of the modular RPs of prokaryotic genomes and metagenomes. It predicts transcription factors (TFs) and two-component systems (TCSs).
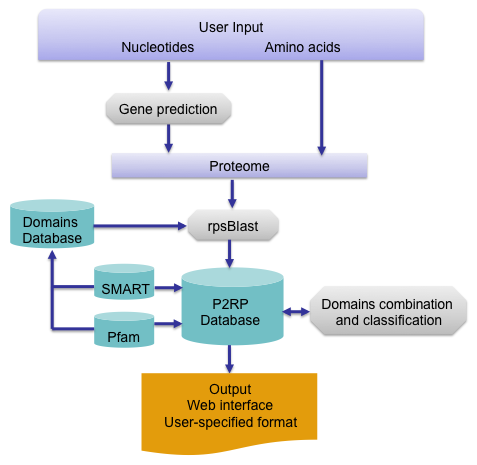
The program can perform several types of analyses depending on the type of request. P2RP requires inputs in Fasta format or gbk file (GenBank) in the case of genome re-annotation. Examples of nucleotide and amino acid sequences are provided to help for testing.
The identification of RP candidates is accomplished by domain analysis of each predicted protein. PHP scripts were developed to search the
numerous combinations of RP modules and to categorise RP proteins into families based on similarity and/or domain architecture. Finally, the cellular
localisation of each RP protein is determined by the presence or absence of transmembrane (TM) segments, using the HMMTOP predictor (Krogh et al., 2001).
The P2RP homepage contains a navigation bar that allows users to execute a new RP analysis, to search for a previous job, to access the help page
or to contact the authors.
The - RP Analysis - menu, allows users to submit one or more sequences for RPs prediction and analysis. The web server takes as input:
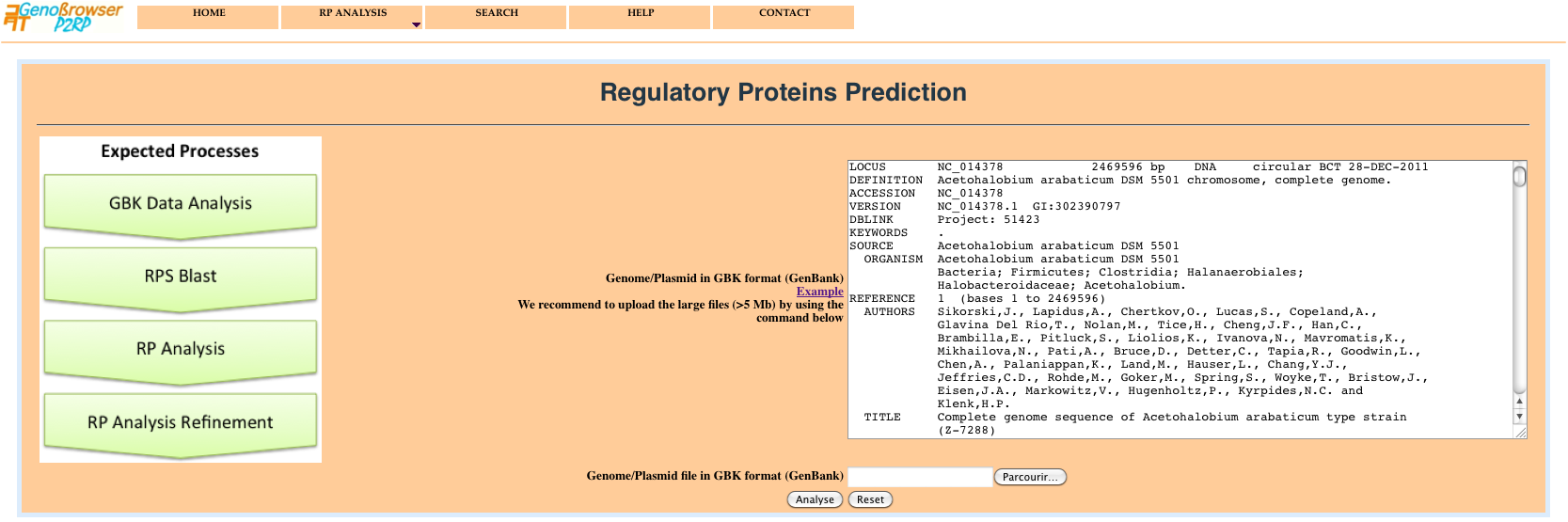
The graphical user interface is provided with a progress bar, used to convey the progress of each task (source data analysis, rpsBlast, RP analysis, RP data refinement). The execution of the different processes is defined by a colour progressive changing using three colours.


Once processing is complete, the results are displayed as a web interface page, and data can be outputted for download in a variety of user-specified formats.
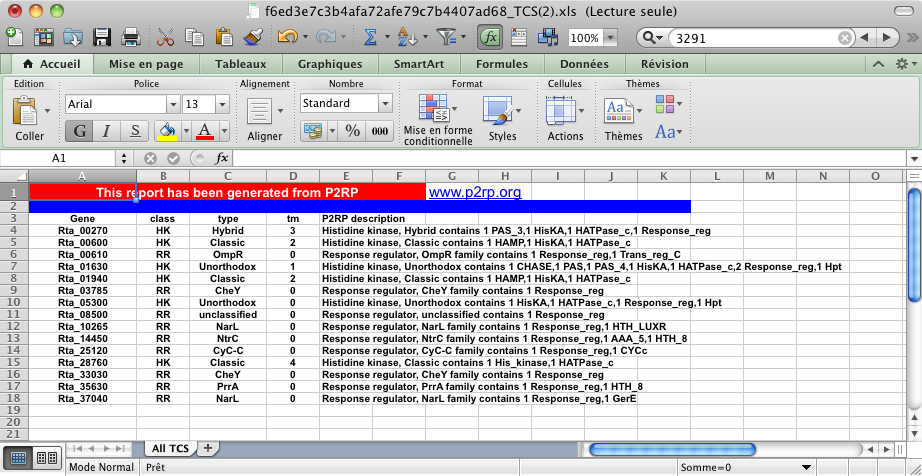
An example of an Excel file output.
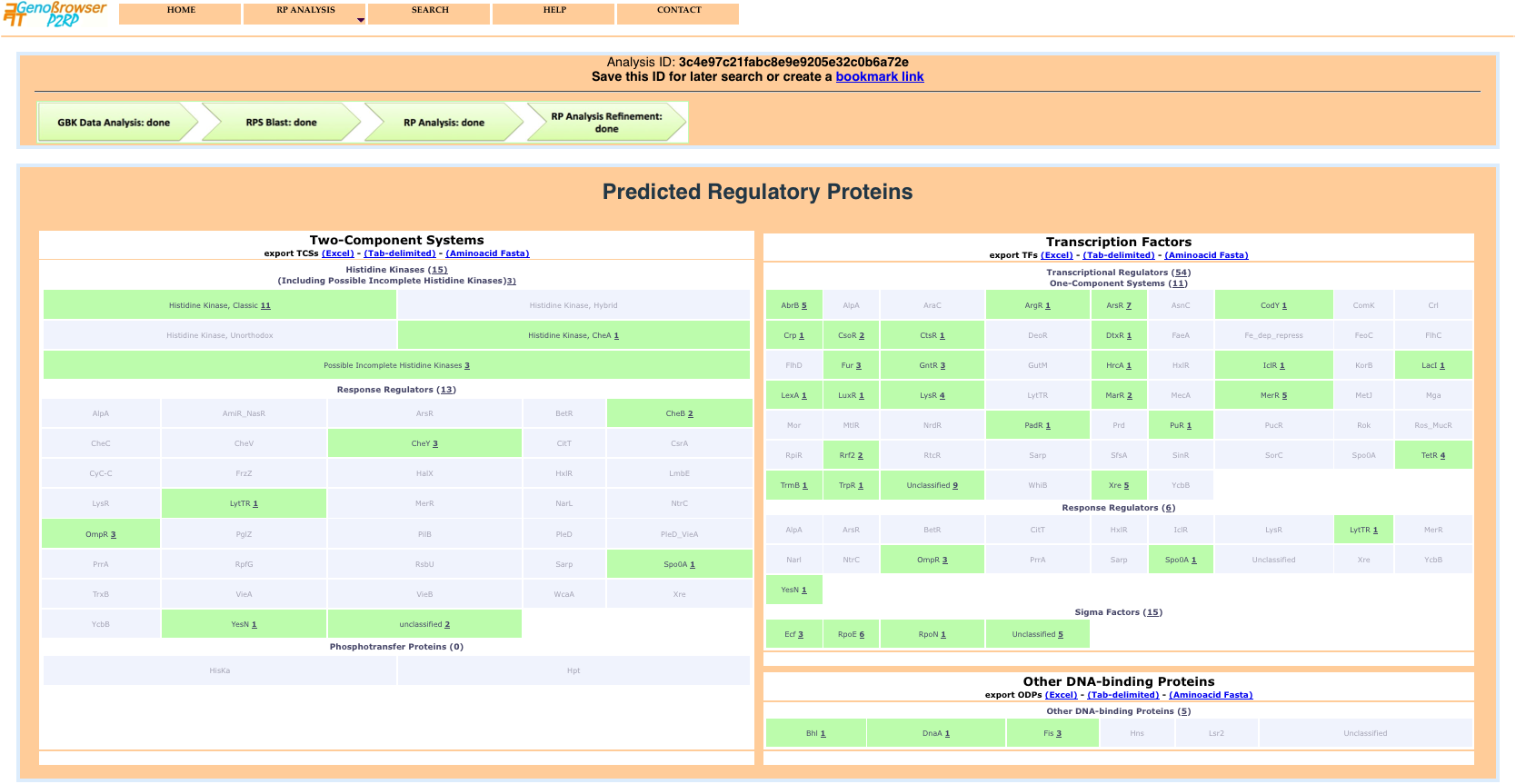
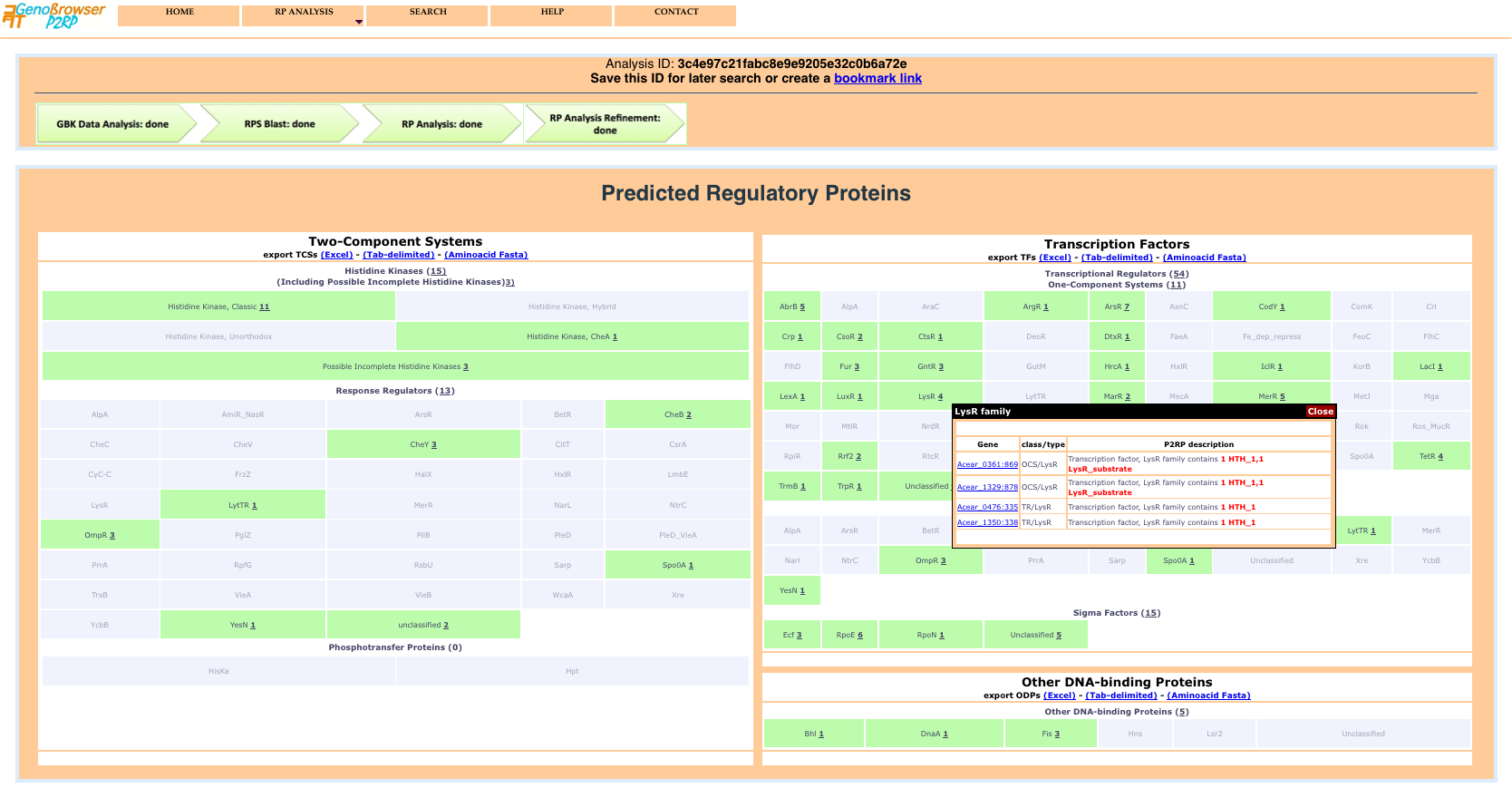
The results page shows global counts of the different categories of RPs and detailed class counts of each category. Each class result provides a detailed gene list, via a popup window, when the mouse is passed over active text.
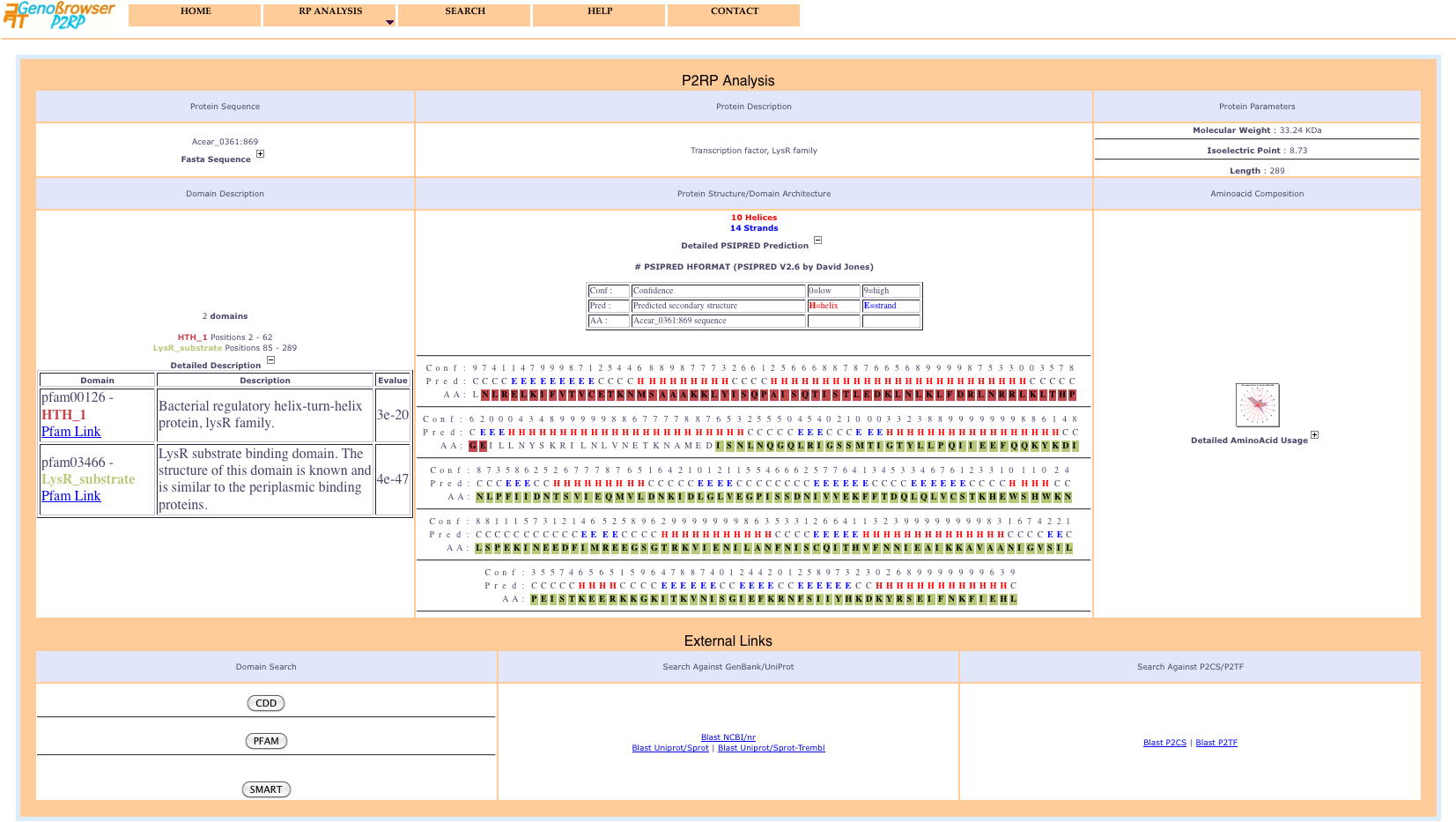
Selecting an object from the identifiers list in the popup window displays a detailed protein description page including 2D structure, domain architecture and aminoacid composition and usage. Blast searches can be performed for the sequence, using external links to numerous public databases.
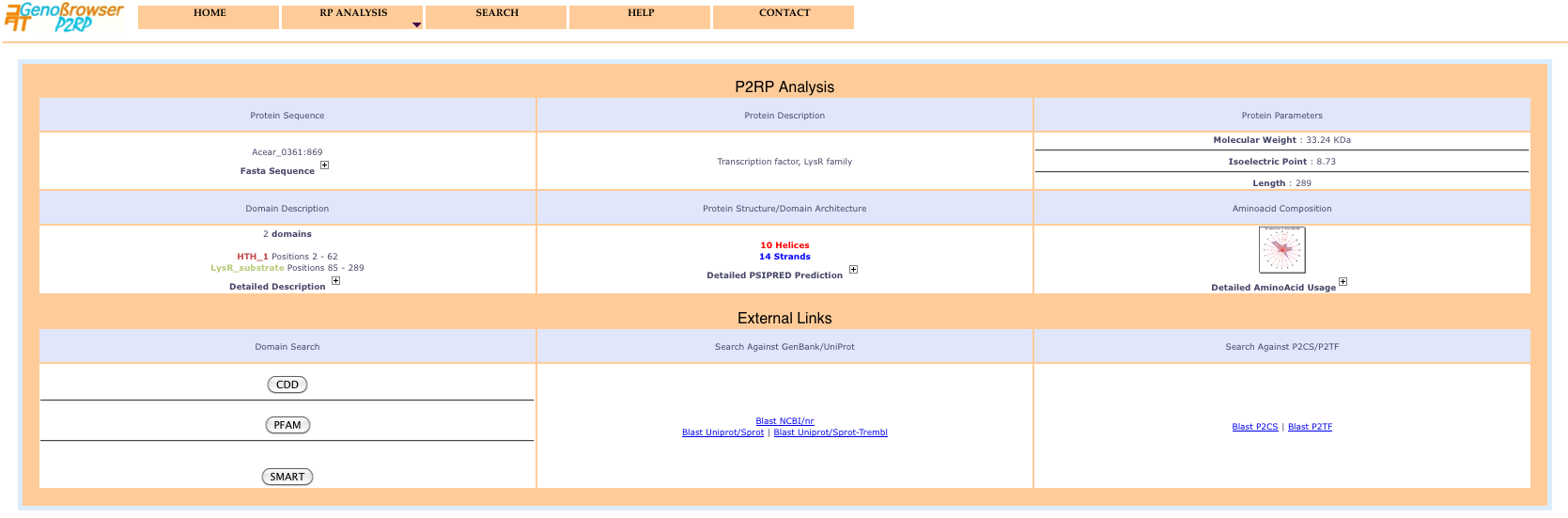
Click on the plus sign (+) to display the corresponding information.
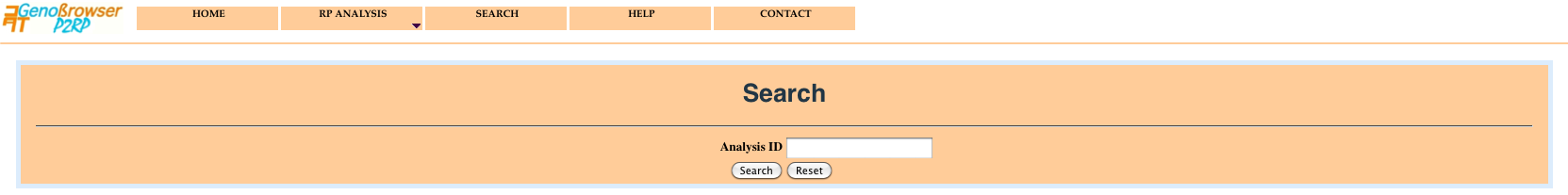
To keep user session private, every user query is given an ID which allows later retrieval of results, using the - Search - menu. So, please remember to save your analysis ID for later use. The data are stored on the server for one month.